Bigelowiella natans Assembly and Gene Annotation
About the Bigelowiella natans genome
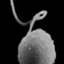
Bigelowiella natans belongs to the eukaryotic 'supergroup' Rhizaria and it is a chlorarachniophyte alga. The chlorarachniophyte alga still possess the nucleomorph, nucleus from the endosymbiotic event in a highly reduced and simplified form. Proteins that function in the plastid of Bigelowiella natans are encoded in 3 separate genomes: the nucleus, the nucleomorph, and the plastid itself. The B.natans genome help in better understanding of the secondary endosymbiotic event.
Assembly
The nuclear genomes of B. natans is approximately 95 megabase pairs (Mb) in size. The genome was assembled by Joint Genome Institute (JGI) into 3736 contigs which are then assembled into 302 scaffolds. It is estimated to have 3.5% sequence bases in the gap.
Annotation
The genome is gene rich with more than 21,000 predicted protein, 229 genes per Mbp scaffold. The average gene length is ~2800nt with average of 8.9 exon per gene. 86% of the genes have intron. 92%of the gene models have start and stop codons.
References
- Algal genomes reveal evolutionary mosaicism and the fate of
nucleomorphs.
Curtis BA, Tanifuji G, Burki F, Gruber A, Irimia M, Maruyama S, Arias MC, Ball SG, Gile GH, Hirakawa Y et al. 2012. Nature. 492:59-65.
Picture credit: Image courtesy of Dr. Geoff McFadden, University of Melbourne, Australia
More information
General information about this species can be found in Wikipedia.
Statistics
Summary
| Assembly | Bigna1, INSDC Assembly GCA_000320545.1, Nov 2012 |
| Database version | 114.1 |
| Golden Path Length | 94,701,163 |
| Genebuild by | JGI |
| Genebuild method | Import |
| Data source | Joint Genome Institute |
Gene counts
| Coding genes | 21,706 |
| Non coding genes | 608 |
| Small non coding genes | 608 |
| Pseudogenes | 6 |
| Gene transcripts | 22,320 |



![Follow us on Twitter! [twitter logo]](/i/twitter.png)
